Question:
Wild-type Drosophila females having three linked genes (AABBCC) were crossed with triple recessive mutant (aabbcc) males. The F1 females (AaBbCc) were backcrossed with aabbcc males. The following F2 progeny numbers were obtained (total = 1000). The gene order is ABC. Find the recombination map distance (in cM) between A and C (round to one decimal place).
\begin{tabular}{l r}
AaBbCc & 241
Aabbcc & 112
aaBbCc & 103
aabbcc & 252
aaBbcc & 17
aabbCc & 134
AabbCc & 14
AaBbcc & 127
\end{tabular}
Wild-type Drosophila females having three linked genes (AABBCC) were crossed with triple recessive mutant (aabbcc) males. The F1 females (AaBbCc) were backcrossed with aabbcc males. The following F2 progeny numbers were obtained (total = 1000). The gene order is ABC. Find the recombination map distance (in cM) between A and C (round to one decimal place).
\begin{tabular}{l r}
AaBbCc & 241
Aabbcc & 112
aaBbCc & 103
aabbcc & 252
aaBbcc & 17
aabbCc & 134
AabbCc & 14
AaBbcc & 127
\end{tabular}
Aabbcc & 112
aaBbCc & 103
aabbcc & 252
aaBbcc & 17
aabbCc & 134
AabbCc & 14
AaBbcc & 127
\end{tabular}
Show Hint
For three-point mapping, {outer-gene distance} (A–C) = SCO\(_{AB}\) + SCO\(_{BC}\) \(+\) \(2\times\)DCO, because each double crossover contributes {two} crossovers between the outer loci.
Updated On: Sep 1, 2025
Show Solution
Verified By Collegedunia
Solution and Explanation
Step 1: Identify nonrecombinants and double crossovers.
The two largest classes are the nonrecombinants (NR) (reflect parental gametes ABC and abc): NR = AaBbCc (241) and aabbcc (252) \(⇒\) \(241+252=493\).
The two smallest classes are the double crossovers (DCO) for the order A–B–C: DCO = AabbCc (AbC) = 14 and aaBbcc (aBc) = 17 \(⇒\) \(14+17=31\).
Step 2: Classify single crossovers (SCO).
For order A–B–C:
\(\bullet\) SCO in A–B interval: Abc (Aabbcc) = 112 and aBC (aaBbCc) = 103 \(⇒\) \(112+103=215\).
\(\bullet\) SCO in B–C interval: ABc (AaBbcc) = 127 and abC (aabbCc) = 134 \(⇒\) \(127+134=261\).
Step 3: Compute A–C map distance.
Map distance is the crossover frequency between the two loci (not just the recombinant frequency of outer markers). Each DCO contains two crossovers between A and C, so DCOs are counted twice: \[ \text{cM}_{A\text{–}C} = \frac{\text{SCO}_{AB} + \text{SCO}_{BC} + 2\times \text{DCO}}{\text{Total}}\times 100 = \frac{215 + 261 + 2\times 31}{1000}\times 100 = \frac{538}{1000}\times 100 = 53.8\ \text{cM}. \]
The two largest classes are the nonrecombinants (NR) (reflect parental gametes ABC and abc): NR = AaBbCc (241) and aabbcc (252) \(⇒\) \(241+252=493\).
The two smallest classes are the double crossovers (DCO) for the order A–B–C: DCO = AabbCc (AbC) = 14 and aaBbcc (aBc) = 17 \(⇒\) \(14+17=31\).
Step 2: Classify single crossovers (SCO).
For order A–B–C:
\(\bullet\) SCO in A–B interval: Abc (Aabbcc) = 112 and aBC (aaBbCc) = 103 \(⇒\) \(112+103=215\).
\(\bullet\) SCO in B–C interval: ABc (AaBbcc) = 127 and abC (aabbCc) = 134 \(⇒\) \(127+134=261\).
Step 3: Compute A–C map distance.
Map distance is the crossover frequency between the two loci (not just the recombinant frequency of outer markers). Each DCO contains two crossovers between A and C, so DCOs are counted twice: \[ \text{cM}_{A\text{–}C} = \frac{\text{SCO}_{AB} + \text{SCO}_{BC} + 2\times \text{DCO}}{\text{Total}}\times 100 = \frac{215 + 261 + 2\times 31}{1000}\times 100 = \frac{538}{1000}\times 100 = 53.8\ \text{cM}. \]
Was this answer helpful?
0
0
Top GATE XL Life Science Questions
- Which one of the following molecules captures \(CO_2\) in the \(CO_4\) cycle?
- Which one of the following methods separates biomolecules based on their hydrodynamic volumes?
- Which one of the following restriction endonucleases is a blunt cutter?
- Which one of the following ion channels opens to repolarize the neuronal membrane when an action potential is generated?
- Which one of the following is the most sensitive immunoassay?
View More Questions
Top GATE XL Life Science Questions
Which one of the given figures P, Q, R, and S represents the graph of the following function? \[ f(x) = |x + 2| - |x - 1| \]

An opaque cylinder (shown below) is suspended in the path of a parallel beam of light, such that its shadow is cast on a screen oriented perpendicular to the direction of the light beam. The cylinder can be reoriented in any direction within the light beam. Under these conditions, which one of the shadows P, Q, R, and S is NOT possible?

- Which one among the following mixtures gives a buffer solution in water?
What is the major product formed in the given reaction?

- The CORRECT order of stability of the given metal oxides is
View More Questions
Top GATE XL Questions
- The CORRECT order of electronegativity is
- Which one of the following is the CORRECT representation of the variation of the Gibbs free energy (G) of a substance with temperature (T) at constant pressure?
- Among the following, the structure representing histidine is
- The CORRECT order of acidity of the following compounds is
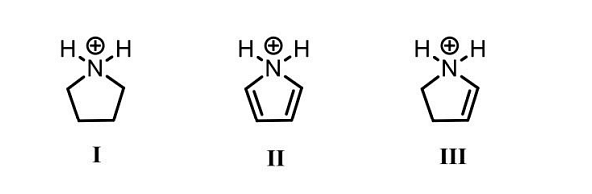
- The molecules A and B are a pair of___
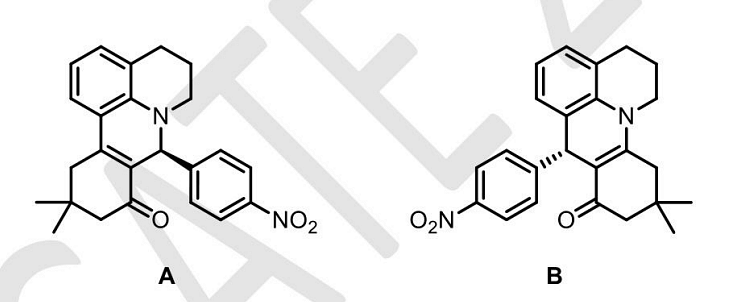
View More Questions